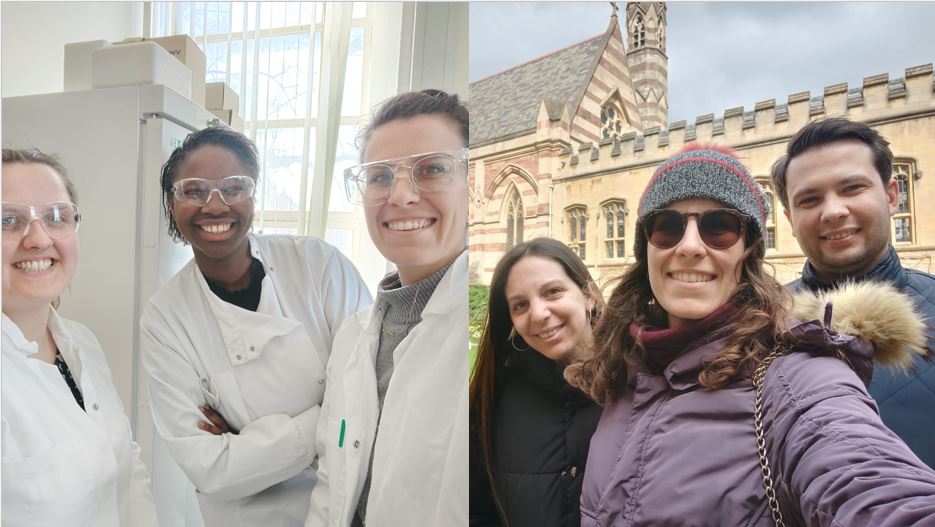
On September 25, the NATURE-ETN ESRs got together for the final project event.
During this online session, the NATURE-ETN ESRs had the chance to hear from alumni of the ClickGene project, a European Training Network in click chemistry and gene therapy, coordinated by Prof. Kellett (DCU) in collaboration with other NATURE-ETN PIs, including Prof. Brown (UOXF), Prof. Carell (LMU), Prof. Hocek (IOCB), and the companies baseclick and ATDBio.
The four ClickGene alumni shared diverse career paths spanning academia, start-ups, and major pharmaceutical companies. Dr. Bastien Viverge (Senior Manager, Regulatory CMC – Clinical Lead, Biogen, CH), Dr. Sarah Walsh (Applied Platform Workflows Manager, Oxford Nanopore Technologies, UK), Dr. Nicolò Zuin Fantoni (Senior Scientist – Oligonucleotides, Roche, CH), and Dr. Georgia Menounou (Postdoctoral Researcher, DCU, IE) each discussed their post-PhD journeys.
We thank the ClickGene alumni for their time and valuable advice to our ESRs.
Dr. Marco Cavallaro (accelCH) also introduced the ESRs to EU funding opportunities for postdocs, industry-academia collaborations, and start-ups/SMEs, highlighting resources that could be useful after obtaining their PhD, whether they pursue careers in academia or industry.